Tag: workflow
Could I re-research my first research?
Last week, one of my teammates, at Old Dominion University, contacted me and asked if she could apply some of the techniques I adopted in the first paper I published during my Ph.D. She asked about the data and any scripts I had used to pre-process the data and implement the analysis. I directed her to where the data was saved along with a detailed explanation of the structure of the directories. It took me awhile to remember where I had saved the data and the scripts I had written for the analysis. At the time, I did not know about data management and the best practices to document my research.
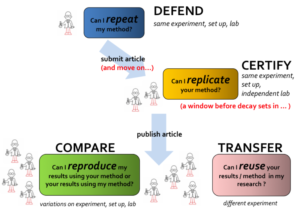
I shared the scripts I generated for pre-processing the data with my colleague, but the information I gave her did not cover all the details regarding my workflow. There were many steps I had done manually for producing the input and the output to and from the pre-processing scripts. Luckily I had generated a separate document that had the steps of the experiments I conducted to generate the graphs and tables in the paper. The document contained details of the research process in the paper along with a clear explanation for the input and the output of each step. When we submit a scientific paper, we get reviews back after a couple of months. That was why I documented everything I had done, so that I could easily regenerate any aspect of my paper if I needed to make any future updates.
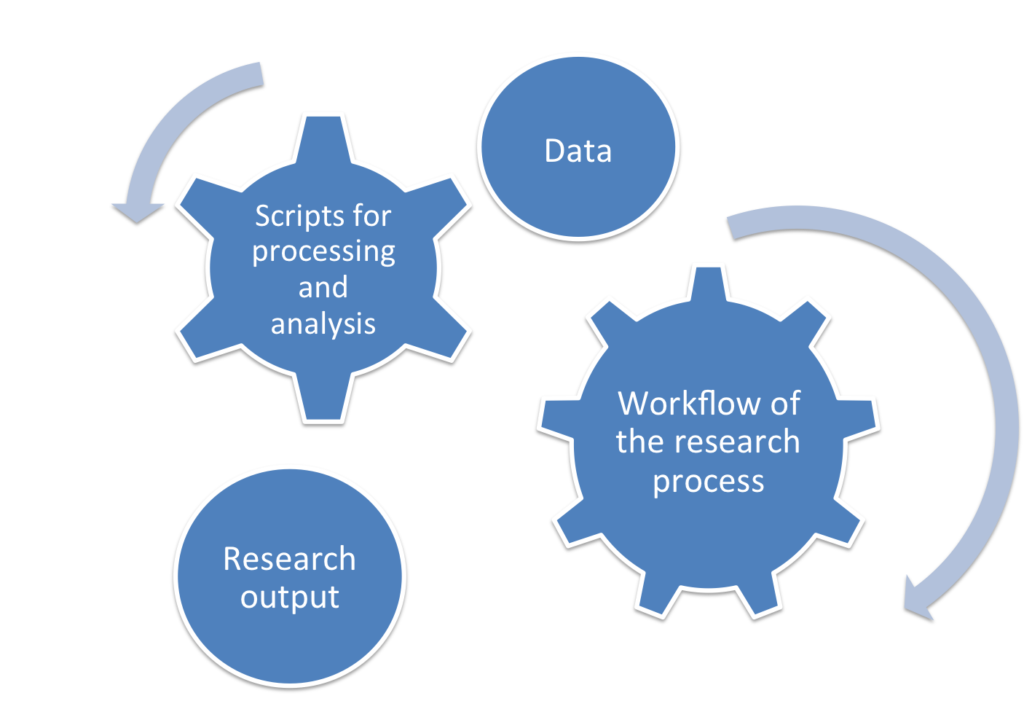
Documenting the workflow and the data of my research paper during the active phase of the research saved me the trouble of trying to remember all the steps I had taken if I needed to make future updates to my research paper. Now my colleague has all the entities of my first research paper: the dataset, the output paper of my research, the scripts that generated this output, and the workflow of the research process (i.e., the steps that were required to produce this output). She can now repeat the pre-processing for the data using my code in a few minutes.
 Funding agencies have data management planning and data sharing mandates. Although this is important to scientific endeavors and research transparency, following good practices in managing research data and documenting the workflow of the research process is just as important. Reproducing the research is not only about storing data. It is also about the best practices to organize this data and document the experimental steps so that the data can be easily re-used and the research can be reproduced. Documenting the directory structure of the data in a file and attaching this file to the experiment directory would have saved me a lot of time. Furthermore, having a clear guidance for the workflow and documentation on how the code was built and run is an important step to making the research reproducible.
Funding agencies have data management planning and data sharing mandates. Although this is important to scientific endeavors and research transparency, following good practices in managing research data and documenting the workflow of the research process is just as important. Reproducing the research is not only about storing data. It is also about the best practices to organize this data and document the experimental steps so that the data can be easily re-used and the research can be reproduced. Documenting the directory structure of the data in a file and attaching this file to the experiment directory would have saved me a lot of time. Furthermore, having a clear guidance for the workflow and documentation on how the code was built and run is an important step to making the research reproducible.

While I was working on my paper, I adopted multiple well known techniques and algorithms for pre-processing the data. Unfortunately, I could not find any source codes that implemented them so I had to write new scripts for old techniques and algorithms. To advance the scientific research, researchers should be able to efficiently build upon past research and it should not be difficult for them to apply the basic tenets of scientific methods. My teammate is not supposed to re-implement the algorithms and the techniques I adopted in my research paper. It is time to change the culture of scientific computing to sustain and ensure the integrity of reproducibility.